Mechanisms of bacterial immunity
Our research group is currently focused on the following topics:
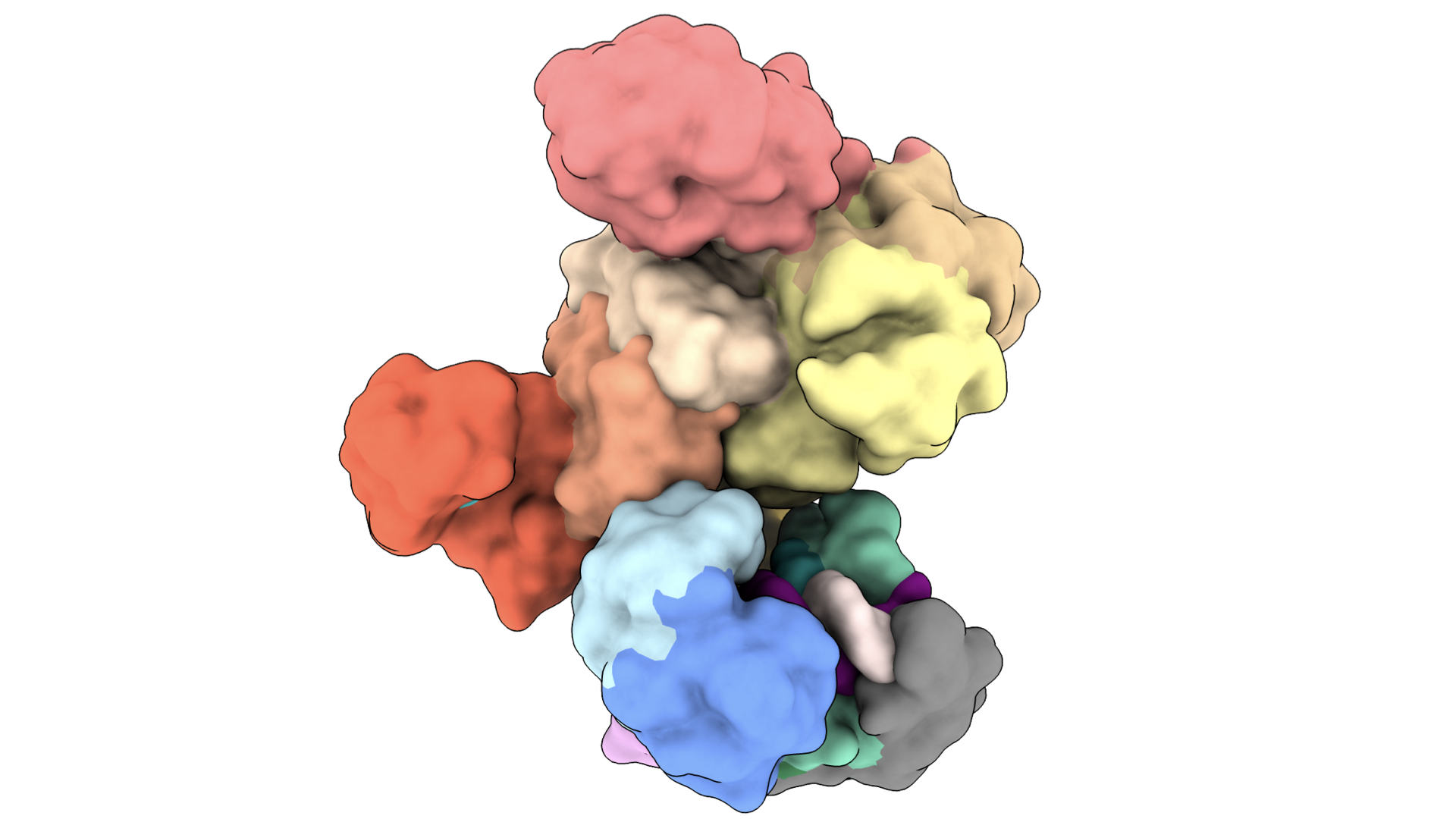
Structures of nucleoprotein complexes
We use a combination of cryo-EM, biochemistry and in vivo assays to understand how immune systems discern self-versus-non-self nucleic acids, and how these systems can be co-opted for novel biotechnology tools. See: Bravo et al., Nature, 2024, Bravo et al., Nature, 2023; Bravo et al., Nature Communications 2022
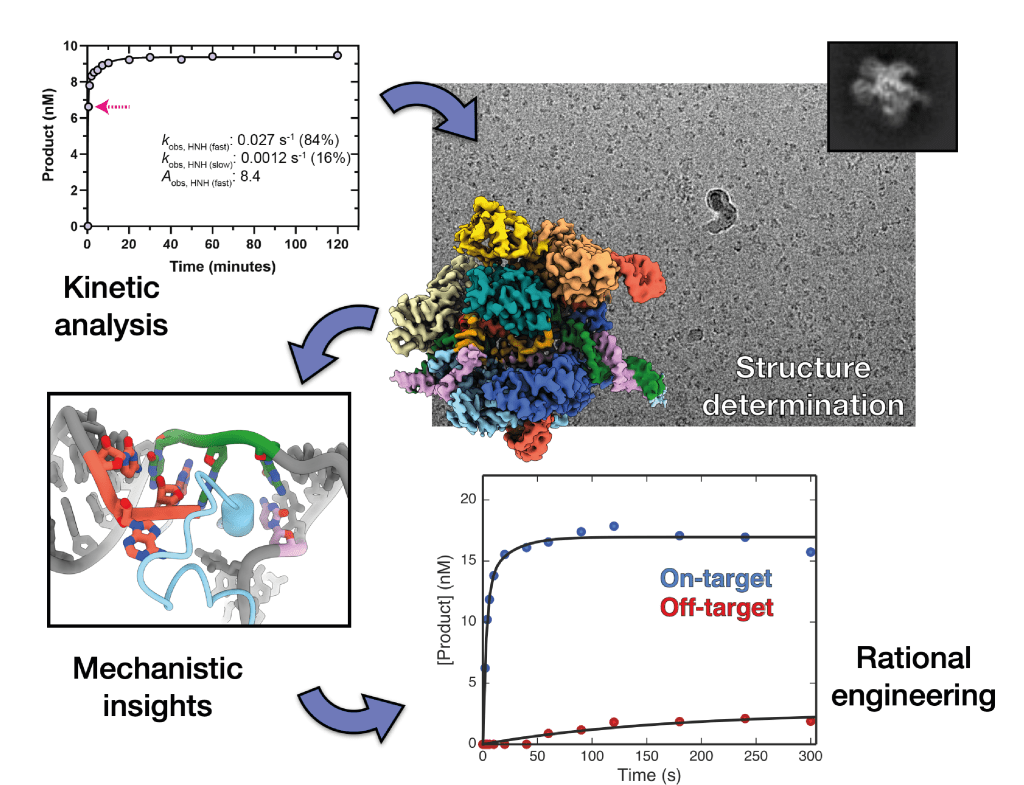
Cryo-EM methods development
Kinetics-guided structural studies to capture transient intermediate states of active complexes. See: Bravo et al., Molecular Cell, 2021; Bravo et al., Nature, 2022; Hibshman et al., Nature Communications, 2024
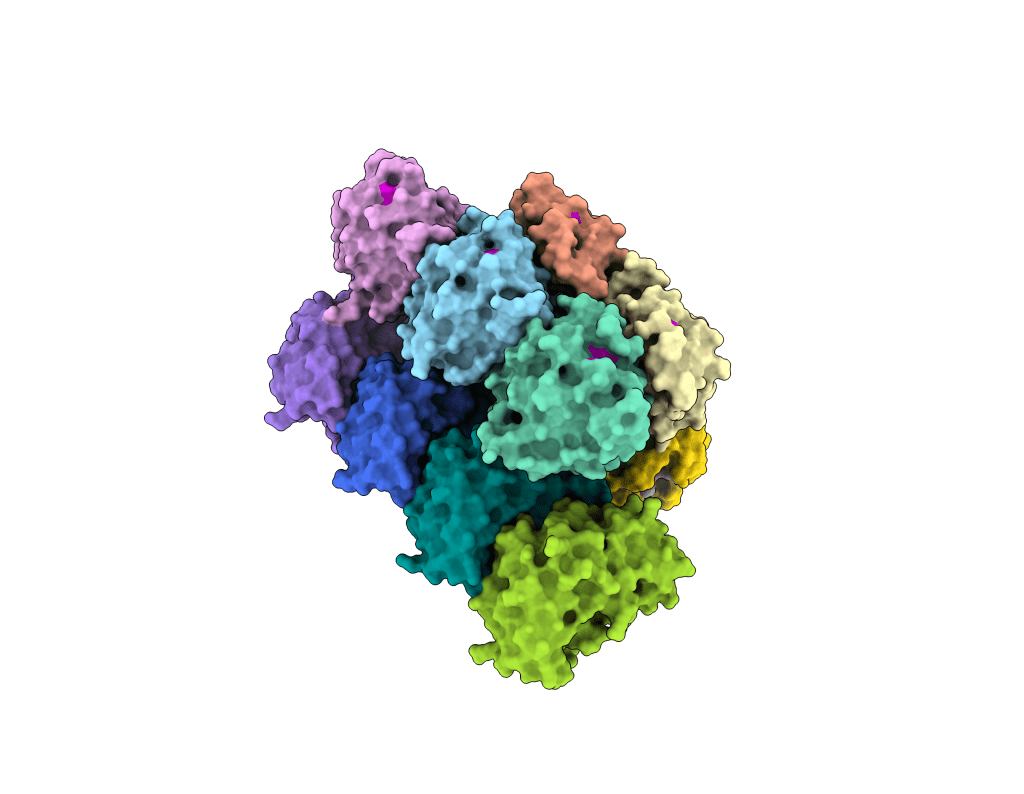
Re-engineering of novel biotechnology tools
We leverage structural and mechanistic insights into bacterial immune systems to engineer precision molecular tools with tailored properties, including enhanced specificity, programmability, and new enzymatic activities suited for genome editing and diagnostics. See: Steens, Bravo & Salazar et al., Science, 2024
Bacteria are major causative agents of disease in humans, and are simultaneously under immense evolutionary pressure from their viral predators — bacteriophages. This dual role has driven the development of numerous sophisticated immune systems providing immunity against invasion. Recent technical breakthroughs have led to the discovery of over 200 previously undetected bacterial immune systems involved in anti-phage defense, antibiotic resistance, and bacterial pathogenesis. While well-known systems such as CRISPR-Cas have been extensively studied, the vast majority of these newly discovered systems remain mechanistically uncharacterized. A central challenge for any immune system is distinguishing self from non-self — a problem that becomes especially acute when the threat arrives as DNA, a molecule the cell must simultaneously protect and surveil. These systems must achieve exquisite specificity to avoid unintended activation and autoimmunity, and many have evolved elaborate mechanisms to gate their activity until a genuine threat is detected.
The Bravo group is particularly interested in understanding how diverse bacterial immune systems identify self from non-self nucleic acids, the structural and mechanistic strategies that enforce this specificity, and the downstream consequences of activation. A recurring theme in our work is how immune systems exploit the physical properties of their targets — such as DNA topology, superhelical tension, or nucleic acid architecture — to achieve discrimination. We are also fascinated by how some systems amplify weak or ambiguous threat signals into decisive immune responses, and how guide molecules are generated, sequestered, and deployed to direct effector activity. We use a combination of structural biology (typically cryo-electron microscopy), biochemistry, biophysics and functional assays to understand the molecular mechanisms that underpin immunity. Our emphasis on cryo-EM and pre-steady state kinetic analysis enables the visualization of transient intermediate conformational states, providing insights into the dynamic and heterogeneous nature of active enzymes.
These novel defense systems represent an untapped treasure trove for precision biotechnology, akin to CRISPR. Our ultimate goal is not only understanding these systems but harnessing that knowledge for rational structure-based engineering. Through this approach, we aim to redesign these defense systems, tailoring them to possess specific, desirable properties. These engineered systems typically exhibit enhanced efficiency and specificity, transforming into potent tools for genome editing and diagnostics. By tapping into the diverse enzymatic activities of newly discovered defense systems, we aspire to unveil a new arsenal of precision tools for molecular biology, poised to shape the future of biotechnological advancements.